Creating value for our customers, suppliers and staff, and ultimately for the public who benefits from scientific discovery.
Faster and Easier Flow Cytometry Sample Prep, Synthetic mRNA for Research and Clinical Labs, Next Generation Cell Recovery for Immuno-Oncology etc.
Anti-Müllerian Hormone (AMH) Quality Control, RIQAS Anti-SARS-CoV-2 Serology EQA Programme, Molecular Controls for Infectious Disease etc.
Tapestri Solution for Tumor Profiling in Single Cell, Ultimate Exonic Coverage, Liquid Biopsy Panels etc.
Direct PCR of Crude Samples without Template Purification, highQu PCR & qPCR Reagents Performance Excellence etc.
Electrospray ionization emitters, RAS proteins for oncology research, MagReSyn NTA beads etc.
Genie Purist: Dedicated for Ultrapure (Type I) Water, Operational Reliability and Ease-of-Use etc.

Tapestri single-cell multiomics analysis can transform how you understand the complexities of multiple myeloma and its precursor stages by integrating genomic, proteomic, and clonotypic subclonal assessment with single-cell resolution, all within a single assay.
The assay includes coverage of key driver and resistance genes, genome-wide copy number variations (CNVs), V(D)J clonotype, and cell lineage & immunotherapy markers.
Webinar: Single-Cell Identification of SMM Clones Driving Myeloma Progression and Therapy Resistance

Based on the core technology Ghost Cytometry which enables supervised or unsupervised machine learning-based sorting, ThinkCyte offers the platform VisionSort.
The system provides cellular fingerprints with morphometric readouts in high resolution, on top of standard fluorescence-based sorting.
VisionSort is the perfect addition to conventional cell sorters, by providing unbiased, label-free cell characterization and sorting, supported by an embedded AI algorithm.

Sample prep made simpler with an automated liquid handling platform that enables cell staining and washing.

RIQAS offers quick report turnaround, ensuring you get your results fast. With a vast user database, you can easily compare data and benchmark your performance. RIQAS is highly accredited, guaranteeing top-notch quality. Enjoy high-quality samples for accurate testing, and benefit from cost-effective, flexible options tailored to your needs. Plus, commutable sample matrices ensure samples mimic the performance of patient samples.
With an comprehensive programme list, RIQAS caters to all your testing needs!

Biolegio offers a range of products, spanning from highly modified and specialised oligos, to standard DNA oligo synthesis.
Applications covered are sequencing, NGS, PCR, Realtime-PCR, SNP detection, genotyping, gene expression and mutation detection.

As the only solution to integrate genotypic and immunophenotypic assessment, the scMRD AML Multiomics Assay targets 40 genes for single-cell DNA sequencing based on current international AML MRD guidelines, such as European LeukemiaNet, and 17-plex antibody-oligonucleotide conjugate (AOC) panel curated for key biomarkers associated with AML MRD.
Through a seamlessly integrated workflow, the assay allows clinician-researchers to:

Superior sensitivity polymerase generating less side products and low copy detection in complex
templates with size up to 6 kb. Highly specific polymerase thanks to efficient blocking at room
temperature. The ALLin™ Buffer includes dNTPs, Magnesium, enhancers, and stabilizers.
10% 3rd Quarter Discount
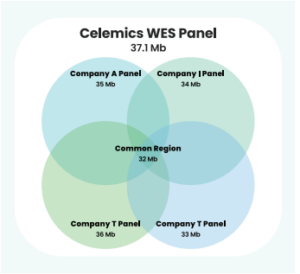
Celemics Whole Exome Sequencing Panel is the most comprehensive WES panel in the market, covering all target regions of major WES panels in the market.
The panel does not compromise uniformity and performs well against hard-to-capture regions such as GC-rich regions.

The chemiSOLO detects low-expressing proteins with femtogram sensitivity and captures marker images at the push of a button.
A unique web browser interface allows the chemiSOLO to be controlled by phone, tablet, or PC, without the need to install any additional software.
In addition to Western blots, the chemiSOLO captures pictures of colorimetric blots or visible stained protein gels, like Coomassie blue or silver stain.
The chemiSOLO’s wide dynamic range can be further enhanced by using EDR, our Extended Dynamic Range function. This feature allows for linear, quantitative data, while avoiding saturation.

The Randox Acusera Serum Indices control is designed to be used to monitor an IVD instrument’s response in the detection of haemolyzed, icteric and lipemic (HIL) samples. This control can be utilised in laboratory interference testing to assist in improving error detection of pre-analytical errors affecting clinical chemistry testing.
This control provides a full range of clinically relevant testing levels, including a negative (-) and three positives (+, ++ & +++)
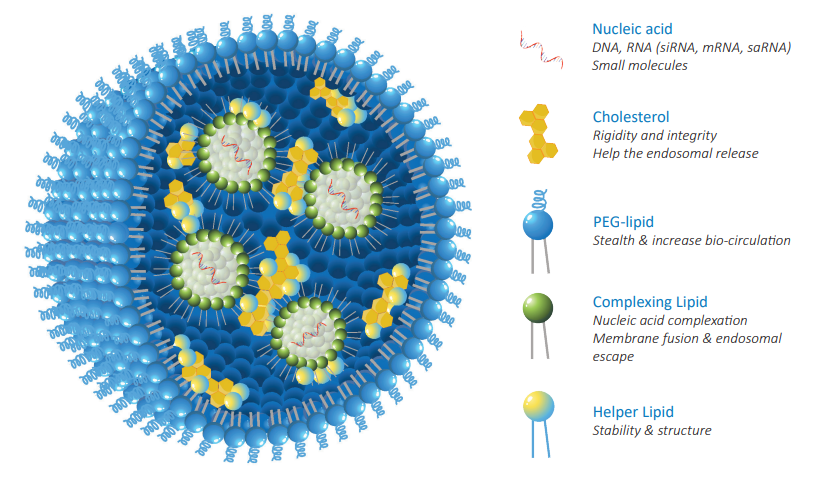
OZ Biosciences can support every stage of your mRNA-LNP production, from mRNA
synthesis to LNP formulation development, manufacturing and fill & finish.
For any of RNA, DNA or APIs encapsulation, you can provide us with your molecule of
interest and we will formulate it into LNP´s.
Monday, Sept 16th, 2024
10:30 – 15.00
Registration – Zoom
Join us for a special seminar where we explore how single-cell DNA multiomics help unveiling cellular complexity. Quantitative characterization of genotype and phenotype at the single-cell level reveals clonal phylogeny and selection at higher resolution, ultimately improving personalized therapeutic selection.
By identifying SNVs, CNVs, and surface protein changes and revealing co-occurrence and zygosity of mutations in single cells, Tapestri enables comprehensive insights into disease progression, development of therapy resistance, and MRD.
Our invited speakers will present data from different areas of research like acute myeloid leukemia (AML), single-cell MRD in AML, functional phenotyping of genomic variants using multiomic scDNA-scRNA-seq, single-cell DNA seq assay for reconstructing the clonal evolution of tumors from FFPE samples, targeted single-cell DNA methylation (scTAM-seq), single-cell Multiple Myeloma multiomics and genome editing.
Tuesday, Sept 10th, 2024
13.00 – 15.00h
Rudbecklaboratoriet, Waldenströmsalen
Join us to discuss news in clonal single cell multiomics analysis with Tapestri. The meeting is arranged in collaboration with the FIMM and will cover:
Enhanced flexibility and cost savings with multiplex capability
Applications: scMRD in AML, sc multiomics for Multiple Myeloma, DNA methylation, genome-wide CNV analysis, gene editing assessment
Open forum discussion
Tuesday, Sept 3rd, 2024
11.00 – 12:30h
FIMM 3rd floor lunch area, Biomedicum 2U
Join us to discuss news in clonal single cell multiomics analysis with Tapestri. The meeting is arranged in collaboration with the FIMM and will cover:
Enhanced flexibility and cost savings with multiplex capability
Applications: scMRD in AML, sc multiomics for Multiple Myeloma, DNA methylation, genome-wide CNV analysis, gene editing assessment
Open forum discussion
Learn how to identify smoldering multiple myeloma (SMM) clonal populations progressing to active myeloma or resistant disease with a true multiomics platform that can quantitatively correlate multiple features from SMM subclones including SNVs, genome-wide CNVs, Ig clonotype, and surface protein expression.
Featuring Adam Sciambi, PhD
CTO, Mission Bio
Thursday June 6 2024, 5:00 pm (CET)
Despite the richer insights that single-cell multi-omics can offer over conventional techniques such as PCR and bulk next-generation sequencing, including characterizing the clonality of heterogeneous cancer cells and changes driving therapy resistance or assessing genome editing outcomes, the costs for sample analysis can be a barrier to adoption for many labs.
To help address this, Mission Bio has recently launched new sample multiplexing features that allow the pooling of multiple samples into a single run on the Tapestri Platform, improving the throughput and productivity, and thereby reducing the per-sample costs for single-cell DNA and protein multi-omic analysis by up to 60%*.
In this educational webinar, we’ll discuss two novel multiplexing approaches available on Tapestri, Antibody Hashing and Genotype Multiplexing. We’ll explore how they enable increased efficiency and versatility while maintaining the same robust performance metrics such as sensitivity and specificity. We’ll also share real-world case studies from labs that are applying multiplexing approaches in disease modeling, CRISPR-editing, and hemato-oncology applications.
We understand our customers’ needs and know how to operate flexibly in a quickly changing but always demanding market.
10 000 +
Products
Instruments and reagents based on innovation for improved laboratory methods.
1 000 +
Satisfied Customers
Networking with 1000´s of individual scientists in Denmark, Finland, Norway, Sweden, Iceland and Baltic countries.
20 +
Years Of Experience
We know how to operate flexibly in a quickly changing but always demanding market.
We are looking for enthusiastic sales professionals who enjoy working in the dynamic life science market.
Menu bar
Contact information
Follow us